Extract a sequence from a construct to create your custom backbone
Extract a backbone from a construct, annotate it and use it quickly in all your designs.
Skip to
Extract sequence for a backbone
Annotate your sequence as a backbone
Upload a donor sequence
Navigate to the toobox and find the plasmid onboarding tool.
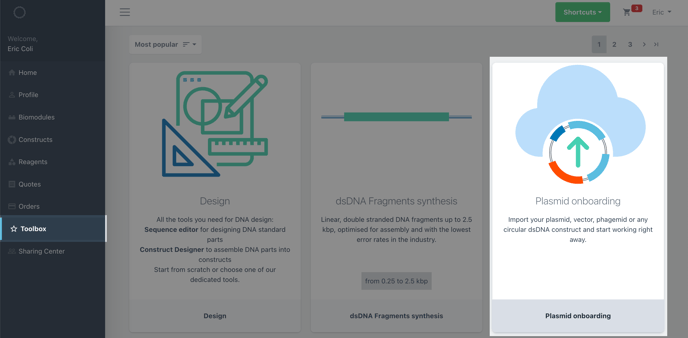
Upload a Genbank or FASTA file of your construct sequence.
Extract your sequence
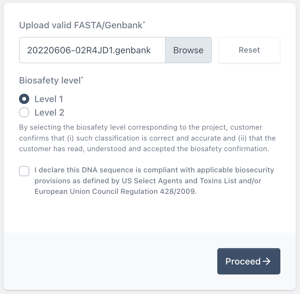
From the sequence viewer, select the region of your sequence that contains your backbone.
You can select regions of the sequence using the select tool (shown below). Alternatively click and drag across your sequence to select that part of the sequence.

Right click on the selected sequence to open the toolkit then choose Extract Sequence.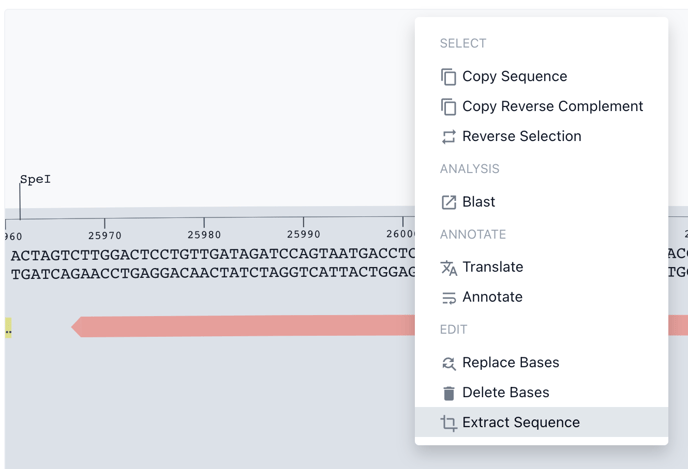
Confirm you would like to Extract Sequence
You have now created a new biomodule containing the sequence you have just extracted.
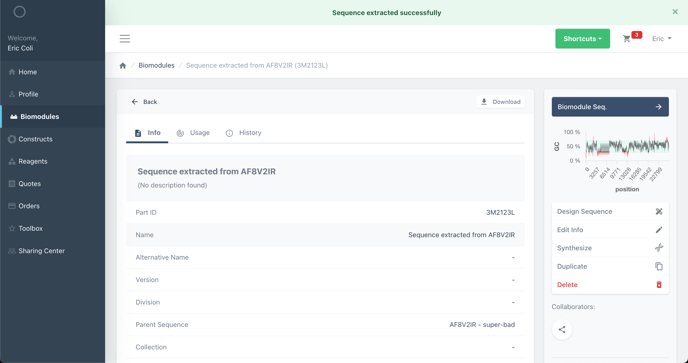
You can extract backbones from both constructs and biomodules, the process is the same. If you choose to upload a linear DNA sequence (biomodule) of only your backbone sequence, you can skip to the next section.
To upload a biomodule containing your backbone see this guide on creating a new biomodule from file.
Annotate your sequence as a backbone
Navigate to your biomodule collection, find the biomodule you just created and hit sequence on the item card.
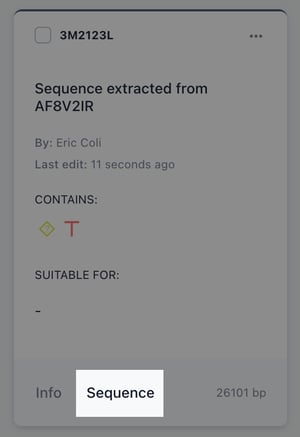
Choose select all to highlight the entire sequence containing your backbone.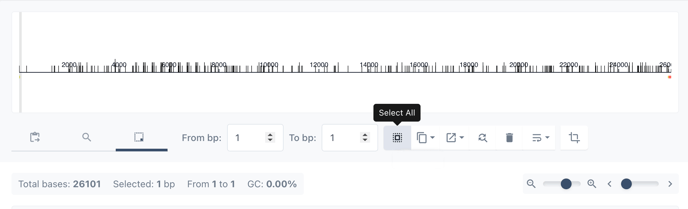
Right click on the highlighted DNA sequence to view the toolkit and choose annotate
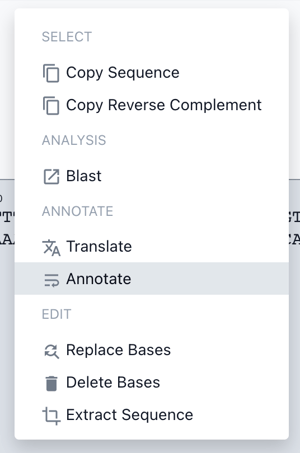
Beneath the annotate as module, search "Vector Backbone" and select that. Give your backbone a name and, optionally, a description. Then Click save.
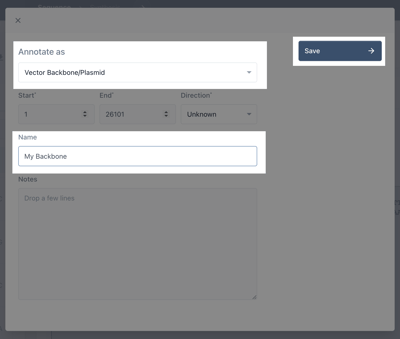
You have now created a backbone annotation.
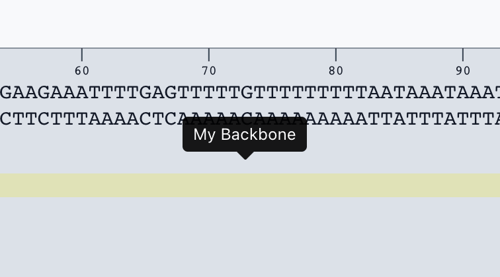
Use your backbone
You can now quickly find your backbone when designing constructs through the advanced search tool.
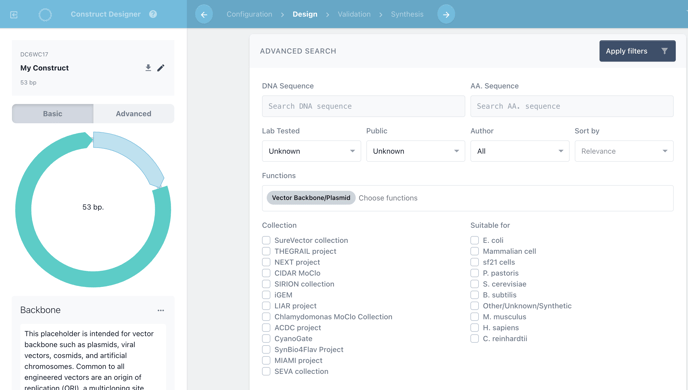
Ordering Cloned Fragments in a custom backbone.
Want to order subcloning in a custom backbone?
See this article on using the backbone you've just created to quickly order clones.